Emerging organoid technologies have been used to model human organ development and diseases “in a dish”. Organoids are derived either from induced pluripotent or organ-restricted adult stem cells. These cells are self-renewing, self-organizing, and able to grow into 3D structures resembling the architecture and physiology of the original biological specimens.
3D organoid models have become indispensable for studying a number of human diseases, including the Zika virus. One of the challenges of organoids is the low reproducibility and high heterogeneity across different cultures and experiments, preventing reliable quantitative comparative analyses between conditions. In addition, current organoid mapping depends mainly on single-cell transcriptome analyses and 2D histology.
To address these problems, Dr. Alexandre Albanese, Dr. Justin Swaney, and colleagues from Dr. Kwanghun Chung’s laboratory (MIT) collaborated with Dr. Paola Arlotta (Harvard University) and Dr. Lee Gehrke (MIT/Harvard Medical School) to perform rapid and multiscale phenotyping of intact cerebral organoids using 3D tissue clearing and imaging (see “Multiscale 3D phenotyping of human cerebral organoids”, Scientific Reports, 2020). Furthermore, this collaboration led to the development of a novel computational pipeline entitled ‘Single-cell and Cytoarchitecture analysis of Organoids using Unbiased Techniques’ (SCOUT).
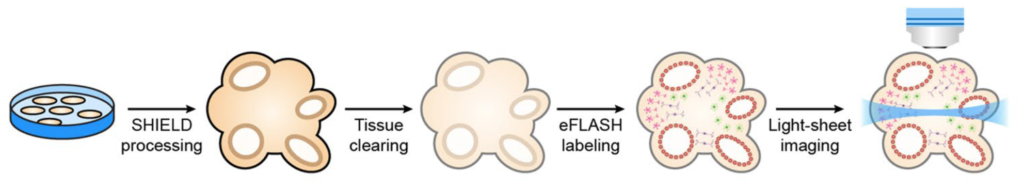
In the Albanese study, induced pluripotent stem cell-derived cerebral organoids were SHIELD-processed and passively cleared using the multipurpose sample-processing device EasyClear.
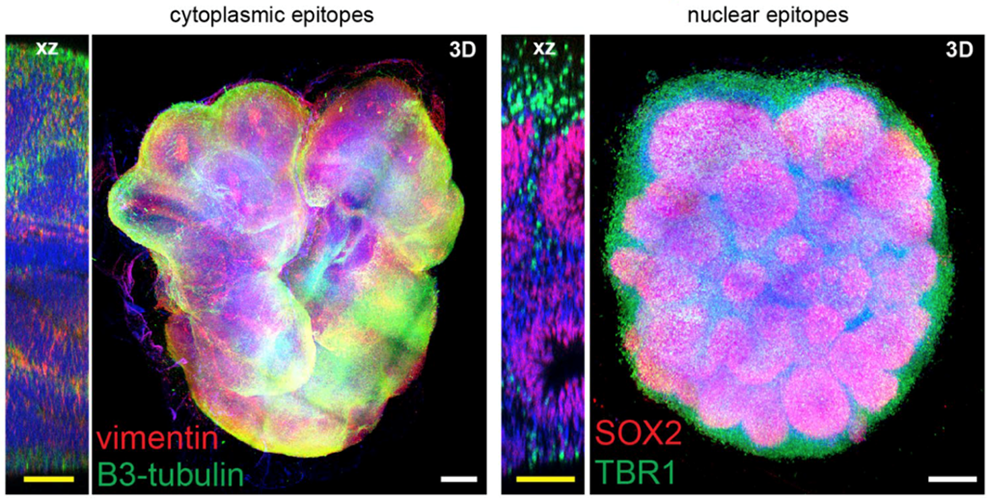
Organoids were then stained for Syto16, SOX2, TBR1, β3-tubulin, MAP2, and vimentin, by using the eFLASH technology developed by the Chung lab. Integrating this technology led to simultaneous staining of 8-10 organoids in only 1-2 days with minimal quantities of antibody required.
Subsequently intact organoids were rapidly imaged (~15 min per organoid), with three-channels at single-cell resolution using the SmartSPIM axially swept light sheet microscope.
This high-throughput workflow from SHIELD, tissue clearing in combination with EasyClear, eFLASH labeling with SmartLabel, and SmartSPIM light sheet imaging resulted in readily achievable and reliable high-resolution 3D data sets which were critical for the development of the SCOUT pipeline.
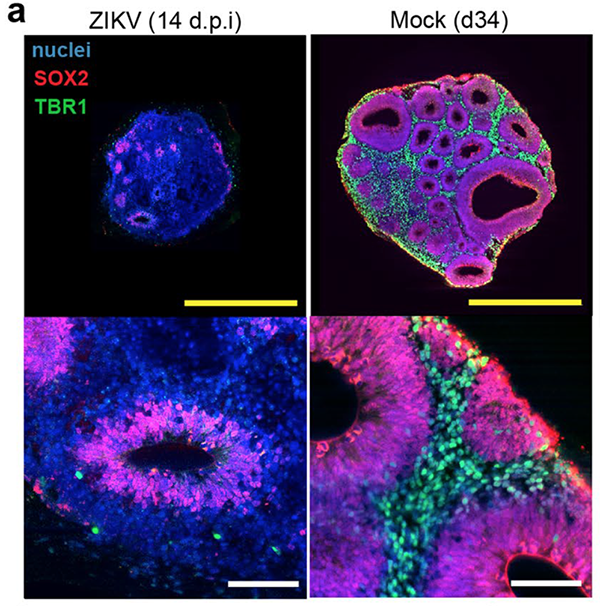
In turn, the SCOUT pipeline enabled automated multiscale comparative 3D analyses of whole organoids and was utilized by Dr. Albanese and colleagues to quantify the impact of Zika virus infection on brain development. The research team showed that Zika infections caused novel reductions in ventricle volume and frequency, and significant differences in the spatial context of rare TBR1+ and SOX2+ cells that are critical for normal brain development.
The use of organoids has significantly accelerated our understanding of human brain development and disease. Implementing LifeCanvas tissue clearing, labeling, and imaging products and devices now enables fast, high-throughput imaging and analyses of these important model systems.
Likewise, other groups have demonstrated the ease of using LifeCanvas products in their organoid research, for example in the lab of Dr. Jennifer A. Lewis at Harvard University, MA (Dr. Skylar-Scott, Dr. Huang, & Dr. Lu et al., 2020) and Dr. Takahashi’s lab at Kyoto University, Japan (Sakaguchi et al., 2019).
Along with the compatibility of LifeCanvas 3D tissue clearing, labeling, and imaging in brain and lung tissues, Dr. Albanese et al. have demonstrated the applicability of this powerful technology for interrogating small and fragile organoids that are critical for furthering our knowledge of neurodevelopment and brain disorders.
Conclusion
- Intact tissue clearing in small, fragile whole organoids
- Simultaneous labeling of at least 10 whole organoids within 1-2 days
- Rapid whole-organoid imaging within 15 minutes