How it started
LifeCanvas Technologies was co-founded by Dr. Kwanghun Chung (Dept. Chemical Engineering, Institute for Medical Engineering and Science, Picower Institute, MIT), one of the pioneers in the field of 3D tissue molecular phenotyping who developed several novel tools to interrogate biological systems (e.g. CLARITY, SWITCH, SE, MAP, SHIELD, eFLASH, ELAST). Using technologies from the Chung lab, LifeCanvas is the only company offering a holistic tissue-processing pipeline from superior sample preservation, passive and active clearing, and fast labeling, to light sheet imaging and analyses. LifeCanvas offers support and guidance through every step of the sample’s journey. In this newsletter, we will provide key insights on the first step of this journey: tissue preservation using SHIELD.
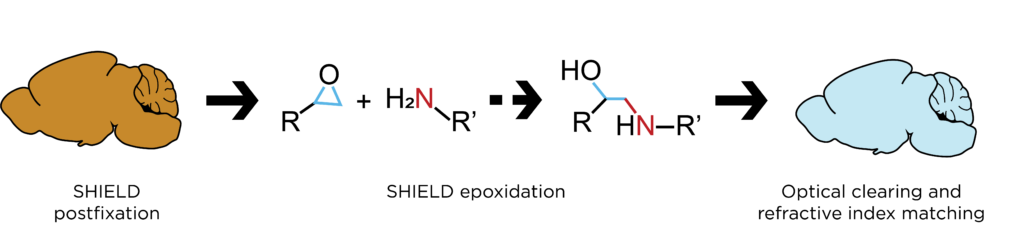
Tissue preservation using SHIELD
SHIELD (Stabilization under Harsh conditions via Intramolecular Epoxide Linkages to prevent Degradation) is a tissue-gel hybridization method using polyfunctional, flexible epoxides to preserve:
- Tissue architecture
- Endogenous fluorescence
- Protein antigenicity
- Nucleic acids
Dr. Young-Gyun Park, Dr. Chang Ho Sohn, and Dr. Ritchie Chen from the Chung lab demonstrated that this polyepoxide forms intra- and intermolecular cross-linkages with biomolecules that enhance the stability of the proteins’ tertiary structure, and preserve the tissue architecture and endogenous fluorescence under harsh chemical conditions (Nature Biotechnology 2018).
This pretreatment is also extremely important in order to proceed to uniform, robust and transparent tissue clearing as well as to complete and uniform immunolabeling.
A major advantage of SHIELD pretreatment is the retention of endogenous fluorescence. SHIELD preserves fluorescent protein signals even under harsh conditions such as heat exposure. For instance, SHIELD-fixed tissues exposed to 70°C for 24 hours exhibited signal retention, compared to dramatically reduced fluorescence in tissues processed only with standard fixatives (paraformaldehyde (PFA) or glutaraldehyde (GA)). Additionally, SHIELD tissues exhibited low tissue autofluorescence, improving the signal to noise ratio in imaged samples.
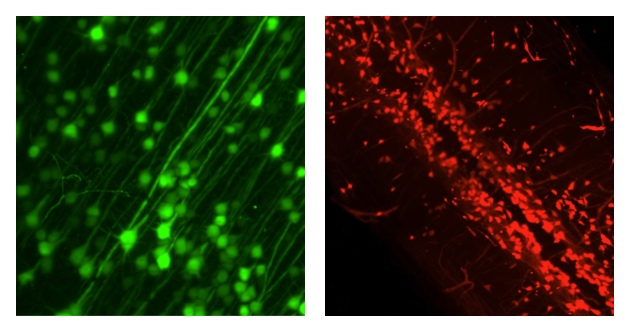
SHIELD’s polyepoxides crosslink proteins and nucleic acids, preserving the probe-binding affinity of antibodies to their epitopes and the binding of ‘fluorescence in situ hybridization’ (FISH) probes to mRNA, while at the same time enabling multiple rounds of labeling. Furthermore, Dr. Park and colleagues showed that SHIELD samples retain tissue integrity with no significant distortion or structural damage after destaining treatments. Preserving tissue architecture is critical for accurate downstream atlas alignment and region segmentation.

SHIELD further enables super-resolution imaging of fine structural and molecular features when combined with MAP (magnified analysis of the proteome). MAP allows the expansion of tissues up to four- to fivefold, resulting in multi-resolution imaging of individual axons and synaptic structures in brain samples.
How to perform tissue preservation with SHIELD in 3 easy steps
Tissue preservation with SHIELD requires the use of three solutions: SHIELD Epoxy solution, SHIELD Buffer solution, and SHIELD ON solution.
Note: The exact amount of solution and incubation times depend on tissue type and size.
Step 1: Perform standard 4% PFA-transcardial perfusion, dissect out tissue of interest, incubate in 4% PFA overnight at 4°C, and store the sample in PBS + 0.02% azide at 4°C.
Step 2: Incubate dissected samples in SHIELD OFF solution at 4°C (mix of SHIELD buffer, SHIELD-epoxy) for 3-4 days.Step 3: Incubate samples in SHIELD ON solution at 37°C (ready-to-use solution). Tissue is ready for clearing!
At this point you can choose to either clear your samples or store them at 4°C for several months. Importantly, long-term storage does not negatively affect fluorescence signals or structural integrity.
Why should I use SHIELD for tissue preservation?
SHIELD tissue preservation is the first essential step in obtaining robust and reliable clearing, labeling, imaging, and analysis results. The SHIELD preservation protocol consists of easy steps and minimal toxic chemical exposure, leading to higher safety and consistency between experiments.
In addition to preventing clearing-induced sample degradation, SHIELD retains tissue stability and probe-binding affinity, allowing multi-round immunolabeling and imaging on the same sample. Furthermore, using SHIELD preservation can save additional costs and labeling time: immunolabeling boosts are typically not needed when using fluorescence reporters or tissues labeled via viral injection, as SHIELD effectively preserves endogenous fluorescence.
Conclusion
SHIELD is one of LifeCanvas Technologies’ reagents among a suite of products geared towards enabling researchers to preserve, clear, label, image, and analyze intact samples. LifeCanvas consists of a multidisciplinary team of experts in neuroscience, cancer, computational biology, machine learning, biomedical engineering, and imaging. Our team strives to be at the forefront of new technological advances, driving research forward with novel reagents, products, and imaging solutions. One of LifeCanvas’ core values is “Integrity and Respect”: as scientists, we understand your needs and goals, and go above and beyond to meet expectations and offer honest advice.
Choosing SHIELD as a tissue preservative enables researchers to implement parts of our fast, streamlined, end-to-end 3D histology workflow: passive clearing using LifeCanvas Passive Clearing Buffer or active clearing and immunolabeling with SmartBatch+, volumetric imaging with SmartSPIM, and analyses with SmartAnalytics. In comparison with other 3D clearing, labeling, and imaging technologies, each step takes days instead of weeks.
Take the first step in using SHIELD tissue preservation for FREE and learn more about our 3D technologies!
Publications citing SHIELD
Swaney et al., 2019; Roy et al., 2019; Martorell et al., 2019; Yun et al., 2019; Benavidez et al., 2020; Muñoz-Castañeda et al., 2020; Ogujiofor et al., 2020; Pop et al., 2020; Foster et al., 2020; Helseth et al., 2021; Luchsinger et al., 2021; Seo et al., 2021; Sweeney et al., 2021; Bharat et al., 2020; Bharat et al., 2021; Albanese et al., 2020


